[2024]
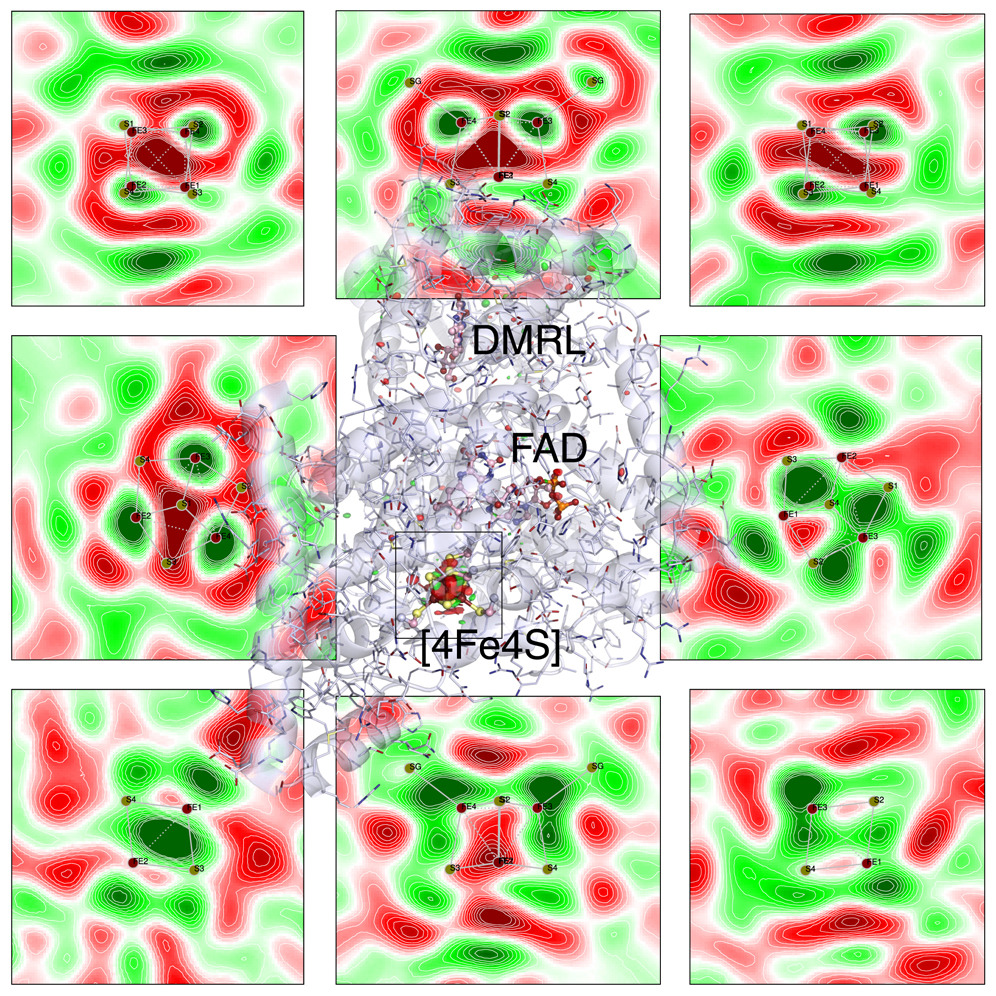
Ren Z* and Yang X*. Deconvolution of dynamic heterogeneity in protein structure. Structural Dynamics. https://doi.org/10.1063/4.0000261 (2024).
Ren Z*, Zhang F, Kang, W, Wang C, Shin H, Zeng X, Gunawardana S, Bowatte K, Kraub N, Lamparter T and Yang X*. Spin-coupled electron densities of iron-sulfur cluster imaged by in situ serial Laue diffraction. Chem https://doi.org/10.1016/j.chempr.2024.02.019 (2024).
Niu K, Wang D, Zhang Y, Biju L, Liu N, Wang X, Ren Z, Lu F*, Yang X* and Zhong D*. Ultrafast Primary Dynamics and Isomerization Mechanism of a Far-Red Sensing Cyanobacteriochrome. J. Phys. Chem. Lett. https://doi.org/10.1021/acs.jpclett.4c00468 (2024).
Kehoe D*, Biswas A, Chen B, Dufour L, Grébert T, Haney A, Joseph KL, Kumarapperuma I, NguyenA, Ratin M, Sanfilippo J, Shukla A, Garczarek L, Yang X, Schluchter WM and Partensky F. Light Color Regulation of Photosynthetic Antennae Biogenesis in Marine Phytoplankton. Plant & Cell Physiology. https://doi.org/10.1093/pcp/pcae115 (2024).
[2023]
Kumarapperuma I., Tom I.P., Bandara S, Montano S and Yang X* Mode of autophosphorylation in bacteriophytochromes RpBphP2 and RpBphP3. Photochem. & Photobiol. Sci. doi:10.1007/s43630-023-00366-9 (2023).
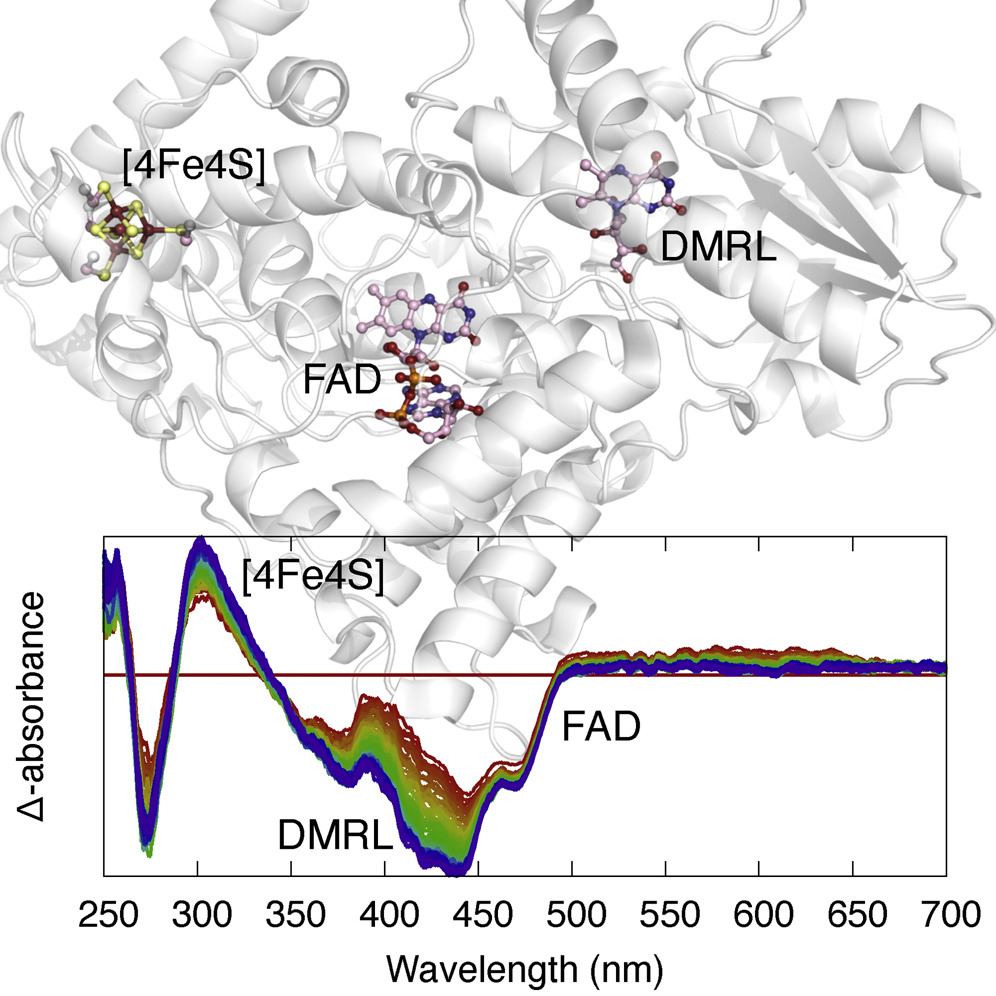
Ren Z*, Kang, W, Gunawardana S, Bowatte K, Thoulass K, Kaeser G, Kraub N, Lamparter T and Yang X*. Dynamic interplays between three redox cofactors in a DNA photolyase revealed by spectral decomposition. Cell Rep. Phys. Sci. doi:10.1016/j.xcrp.2023.101297 (2023).
[2022]
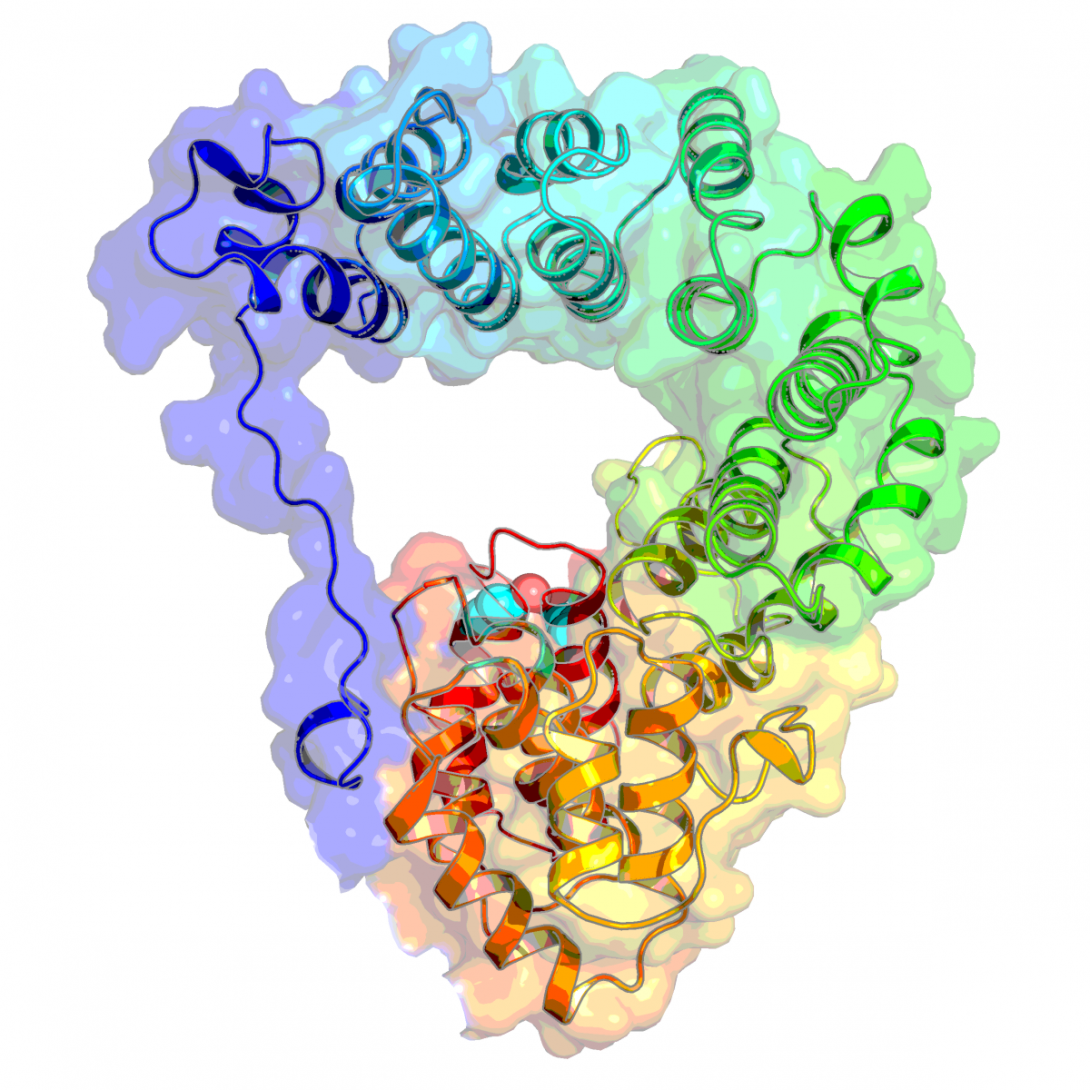
Kumarapperuma I, Joseph K-L, Wang C, Biju LB, Tom IP, Weaver KD, Grebert T, Partensky F, Schluchter MW*, and Yang X* Crystal structure and molecular mechanism of an E/F type bilin lyase-isomerase. Structure. DOI:https://doi.org/10.1016/j.str.2022.01.007. (2022). A commentary by Maayan Suissa-Szlejf, Jenia Sklyar, and Noam Adir. Who’s your neighbor? Suggesting alternative assemblies of E/F type bilin lyases from crystal lattice analysis. Structure 30, 534-536 https://doi.org/10.1016/j.str.2022.03.015 (2022).
Biju L. M., Wang C, Kang W, Tom I. P. , Kumarapperuma I, Yang X* and Ren Z* On-chip Crystallization and Large-Scale Serial Diffraction at Room Temperature. JoVE. doi: 10.3791/63022 (2022).
Lee SJ, Kim TW, Kim JG, Yang C, Yun SR, Kim C, Ren Z, Kumarapperuma I, Kuk J, Moffat K, Yang X* and Ihee H* Light-induced protein structural dynamics in bacteriophytochrome revealed by time-resolved x-ray solution scattering. Science Advances. (2022).
Ren, Z. Photoinduced Isomerization Sampling of Retinal in Bacteriorhodopsin. PNAS Nexus (2022). (The preprint was published on BioRxiv doi: https://doi.org/10.1101/2021.09.16.460656, 2021).
Carrigee L, Frick JP, Liu X, Karty J, Trinidad J, Tom IP, Yang X, Dufour L, Partensky F and Schluchter WM The phycoerythrobilin isomerization activity of MpeV in Synechococcus sp. WH8020 is prevented by the presence of a histidine at position 141 within its phycoerythrin-I β-subunit substrate. Front. Microbiol. https://doi.org/10.3389/fmicb.2022.1011189 (2022).
[2021]
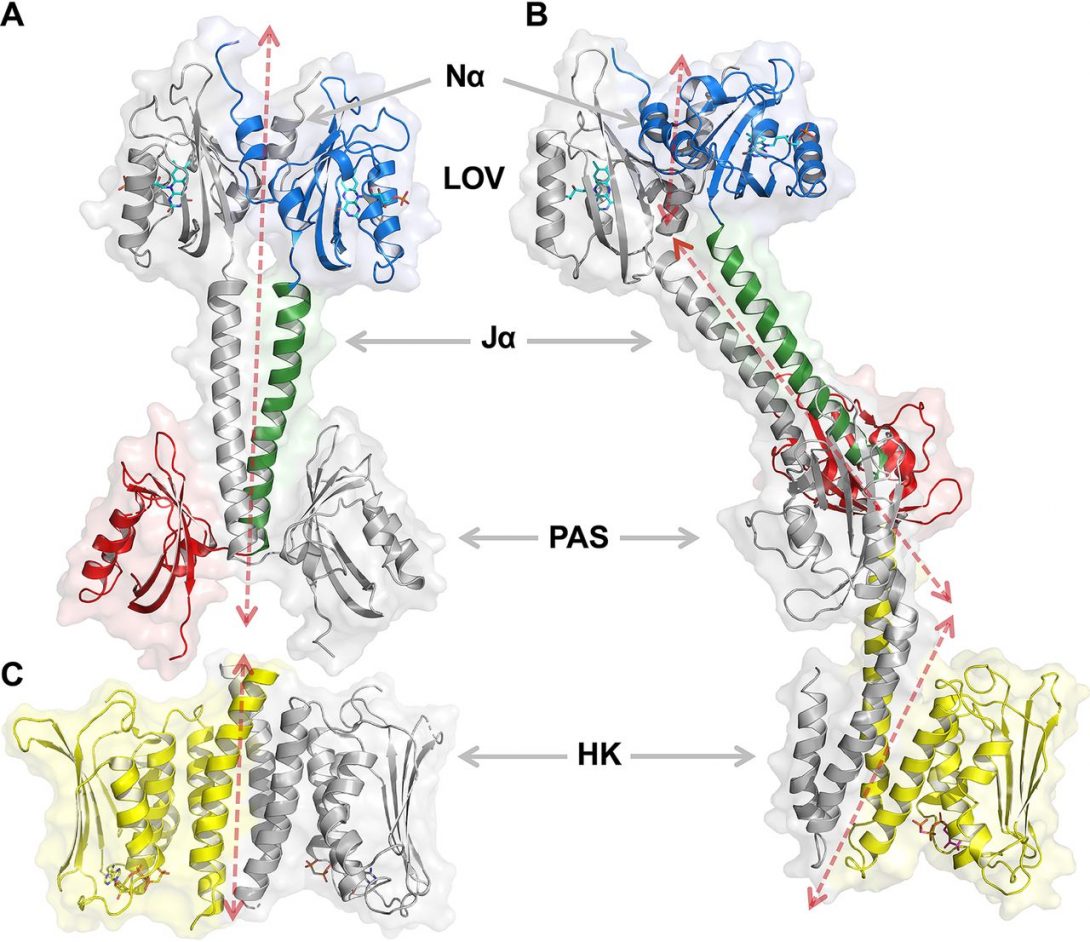
Rinaldi J#, Fernández I#, Shin H#, Sycz G, Gunawardana S, Kumarapperuma I, Paz JM, Otero LH, Cerutti ML, Á Zorreguieta Á, Ren Z, Klinke S*, Yang X*, Goldbaum FA*. Dimer Asymmetry and Light Activation Mechanism in Brucella Blue-Light Sensor Histidine Kinase. mBio https://doi.org/10.1128/mBio.00264-21 (2021).
Bandara S, Rockwell N, Zeng X, Ren Z, Wang C, Shin H, Martin SS, Moreno MV, Lagarias JC* & Yang X* Crystal Structure of a far-red-sensing cyanobacteriochrome reveals an atypical bilin conformation and spectral tuning mechanism. PNAS doi:10.1073/pnas.2025094118 (2021)
Ren, Z. Directional Proton Conductance in Bacteriorhodopsin Is Driven by Concentration Gradient, Not Affinity Gradient. BioRxiv doi: https://doi.org/10.1101/2021.10.04.463074 (2021).
[2020]
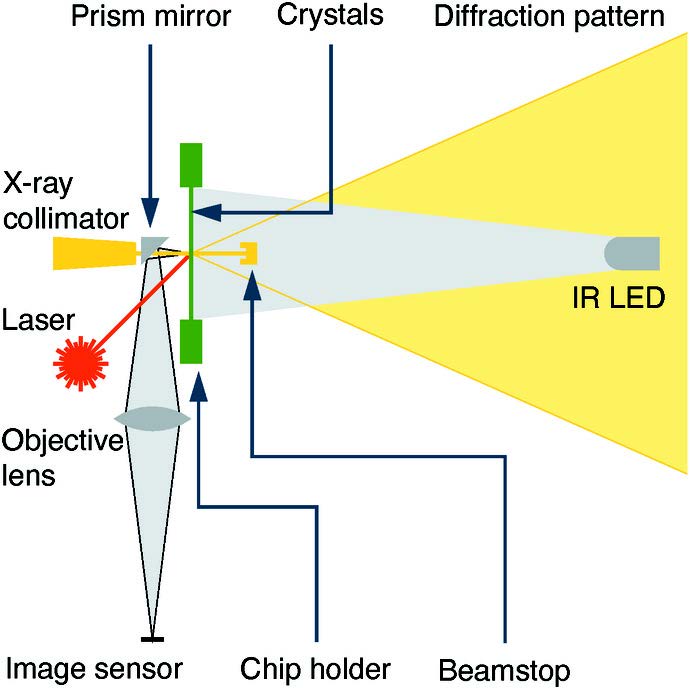
Ren Z*, Wang C, Shin H, Bandara S, Kumarapperuma I, Ren MY, Kang W & Yang X* An automated platform for in situ serial crystallography at room temperature IUCrJ 7:1009 doi:10.1107/S2052252520011288 (2020)
Wang D, Li X, Zhang S, Wang L, Yang X & Zhong D* Revealing the origin of multiphasic dynamic behaviors in cyanobacteriochrome PNAS 117:19731 doi:10.1073/pnas.2001114117 (2020)
Wang D, Li X, Wang L, Yang X* & Zhong D* Elucidating Ultrafast Multiphasic Dynamics in the Photoisomerization of Cyanobacteriochrome J Phys Chem Lett 11:8819-8824 doi:10.1021/acs.jpclett.0c02467 (2020)
Wang D, Qin Y, Zhang M, Li X, Wang L, Yang X & Zhong D* The Origin of ultrafast multiphasic dynamics in photoisomerization of bacteriophytochrome J Phys Chem Lett 11:5913 doi:10.1021/acs.jpclett.0c01394 (2020)
Slavov C, Fischer T, Barnoy A, Shin H, Rao AG, Wiebeler C, Zeng X, Sun Y, Xu Q, Gutt A, Zhao K-H, Gärtner W, Yang X*, Schapiro I* & Wachtveitl J* The interplay between chromophore and protein determines the extended excited state dynamics in a single-domain phytochrome PNAS 117:16356 doi:10.1073/pnas.1921706117 (2020)
[2019]
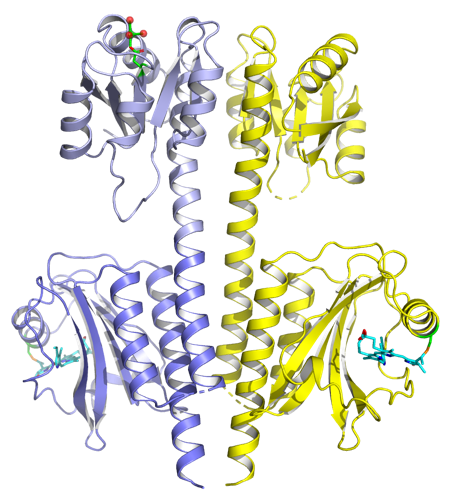
Ren Z* Ultrafast structural changes decomposed from serial crystallographic data J Phys Chem Lett 10:7148 doi:10.1021/acs.jpclett.9b02375 (2019)
Wang D, Qin Y, Zhang S, Wang L, Yang X & Zhong D* Elucidating the molecular mechanism of ultrafast Pfr-state photoisomerization in bathy bacteriophytochrome PaBphP J Phys Chem Lett 10:6197 doi:10.1021/acs.jpclett.9b02446 (2019)
Shin H, Ren Z, Zeng X, Bandara S & Yang X* Structural basis of molecular logic OR in a dual-sensor histidine kinase PNAS 116:19973 doi:10.1073/pnas.1910855116 (2019)
Wang C, Ralko A, Ren Z, Rosenhouse-Dantsker A & Yang X* Modes of cholesterol binding in membrane proteins: A joint analysis of 73 crystal structures In: Rosenhouse-Dantsker A, Bukiya A (eds) Direct mechanisms in cholesterol modulation of protein function. Advances in Experimental Medicine and Biology 1135:67 Springer, Cham doi:10.1007/978-3-030-14265-0_4 (2019)
[2017-2018]
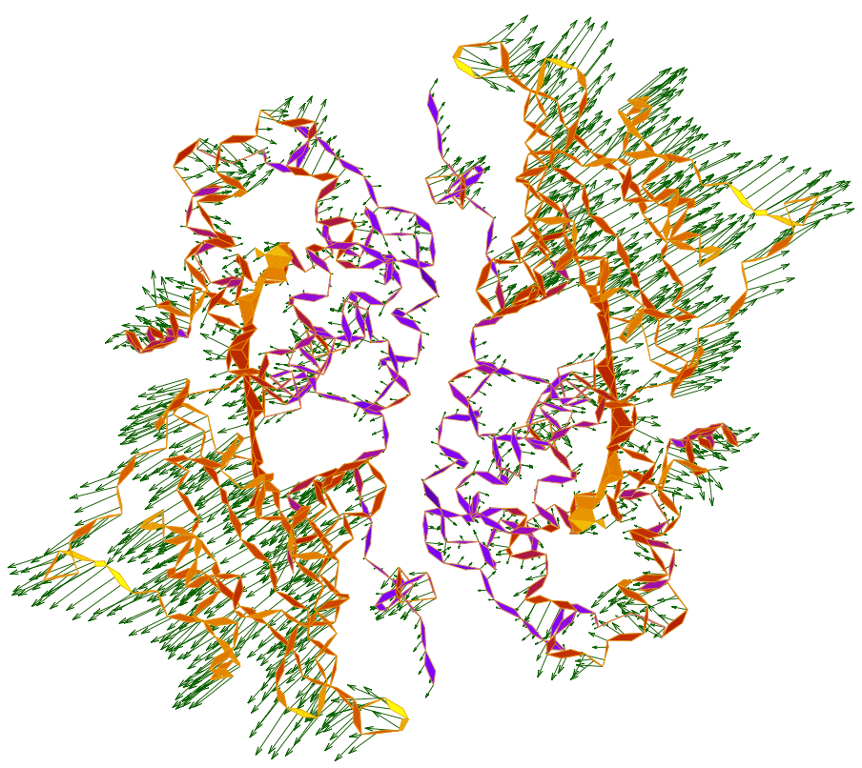
Ren Z*, Ayhan M, Bandara S, Bowatte K, Kumarapperuma I, Gunawardana S, Shin H, Wang C, Zeng X & Yang X* Crystal-on-crystal chips for in situ serial diffraction at room temperature Lab Chip 18:2246 doi:10.1039/C8LC00489G (2018)
Ren Z Single crystal quartz chips for protein crystallization and X-ray diffraction data collection and related methods US patent 9632042 http://patft.uspto.gov/netacgi/nph-Parser?Sect1=PTO1&Sect2=HITOFF&d=PALL&p=1&u=%2Fnetahtml%2FPTO%2Fsrchnum.htm&r=1&f=G&l=50&s1=9632042.PN.&OS=PN/9632042&RS=PN/9632042 (2017)
Bandara S, Ren Z*, Lu L, Zeng X, Shin H, Zhao K-H* & Yang X* Photoactivation mechanism of a carotenoid-based photoreceptor PNAS 114:6286 doi:10.1073/pnas.1700956114 (2017)
Kim TH, Mehrabi P, Ren Z, Sljoka A, Ing C, Bezginov A, Ye L, Pomès R, Prosser RS* & Pai EF* The role of dimer asymmetry and protomer dynamics in enzyme catalysis Science 355:262 doi:10.1126/science.aag2355 (2017) [Commentary by Saleh T & Kalodimos CG Enzymes at work are enzymes in motion Science 355:274 doi:10.1126/science.aal4632 (2017)]
Zhang F, Ma H, Kwiatkowski D, Bowatte K, Mittmann E, Qasem H, Krauss N, Zeng X, Ren Z, Scheer P*, Yang X* & Lamparter T* Crystal structures of bacterial (6-4) photolyase mutants with impaired DNA repair activity Photochem Photobiol 93:304 doi:10.1111/php.12699 (2017)
[2016]
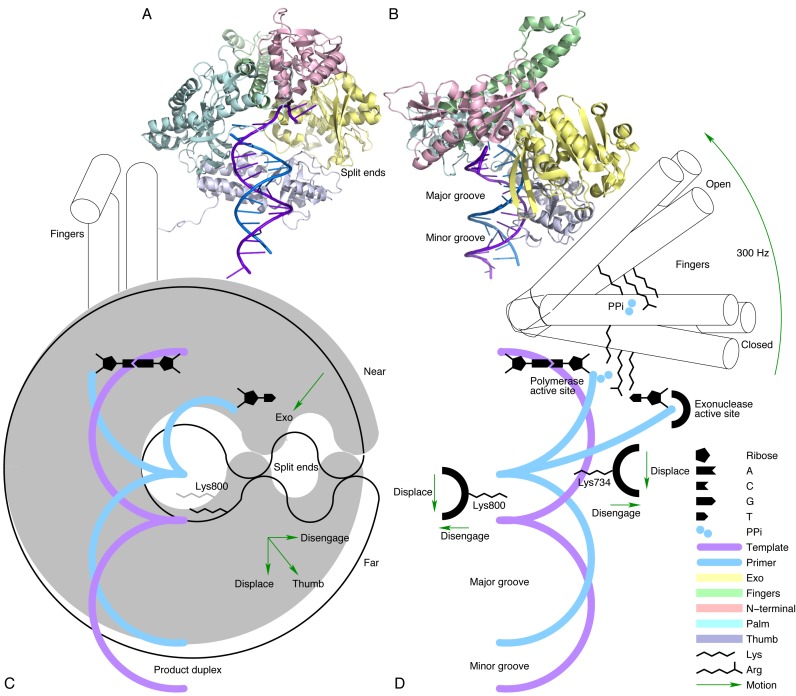
Wang C, Flanagan ML, McGillicuddy RD, Zheng H, Ginzburg AR, Yang X, Moffat K & Engel GS Bacteriophytochrome photoisomerization proceeds homogeneously despite heterogeneity in ground state Biophys J 111:2125 doi:10.1016/j.bpj.2016.10.017 (2016)
Ren Z*, Ren PX, Basulu R & Yang X* Transmembrane helices tilt, bend, slide, torque, and unwind between functional states of rhodopsin Sci Rep 6:34129 doi:10.1038/srep34129 (2016)
Ren Z* Molecular events during translocation and proofreading extracted from 200 static structures of DNA polymerase Nucleic Acid Research 44:7457 doi:10.1093/nar/gkw555 (2016)
Ren Z* & Yang X Angular-split/temporal-delay approach to ultrafast protein dynamics at XFELs Acta Cryst D 72:871 doi:10.1107/S2059798316008573 (2016)
[2015]
Yang X*, Montano S & Ren Z How does photoreceptor UVR8 perceive a UV-B signal? Photochem Photobiol 91:993 doi:10.1111/php.12470 (2015)
Pawate AS, Srajer V, Schieferstein J, Guha S, Henning R, Kosheleva I, Schmidt M, Ren Z, Kenis PJA & Perry SL* Towards time-resolved serial crystallography in a microfluidic device Acta Cryst F 71:823 doi:10.1107/S2053230X15009061 (2015)
Yang X*, Stojković EA, Ozarowski WB, Kuk J, Davydova E & Moffat K* Light signaling mechanism of two tandem bacteriophytochromes Structure 23:1179 doi:10.1016/j.str.2015.04.022 (2015)
Yang XF, Zeng X, Moffat K & Yang X* Structure of the response regulator RPA3017 involved in red-light signaling in Rhodopseudomonas palustris Acta Cryst F 71:1215 doi:10.1107/S2053230X15014661 (2015)
Zeng X, Ren Z, Wu Q, Fan J, Peng PP, Tang K, Zhang R, Zhao KH & Yang X* Dynamic crystallography reveals early signaling events in ultraviolet photoreceptor UVR8 Nature Plants 1:14006, doi:10.1038/nplants.2014.6 (2015)
Wu Q, Huang B, Niehaus TA, Yang X, Fan J* & Zhang RQ* The role of tryptophans in the UV-B absorption of a UVR8 photoreceptor – a computational study Phys Chem Chem Phys 17:10786 doi:10.1039/C4CP06073C (2015)
Scheer H*, Yang X & Zhao KH* Biliproteins and their applications in bioimaging Procedia Chem 14:176 doi:10.1016/j.proche.2015.03.026 (2015)
[2014]
Perry SL*, Guha S, Pawate AS, Henning R, Kosheleva I, Srajer V, Kenis PJA & Ren Z* In situ serial Laue diffraction on a microfluidic crystallization device J Appl Cryst 47:1975, doi:10.1107/S1600576714023322 (2014)
Zhou W, Ding WL, Zeng X, Dong LL, Zhao B, Zhou M, Scheer H, Zhao KH* & Yang X* Structure and mechanism of the phycobiliprotein lyase CpcT J Biol Chem 289:26677, doi:10.1074/jbc.M114.586743 (2014)
Van Thor JJ*, Warren MM, Lincoln CN, Chollet M, Lemke HT, Fritz DM, Schmidt M, Tenboer J, Ren Z, Srajer S, Moffat K & Graber T Signal to noise considerations for single crystal femtosecond time resolved crystallography of the photoactive yellow protein Faraday Discuss 171:439, doi:10.1039/c4fd00011k (2014)
Peng PP, Dong LL, Sun YF, Zeng X, Ding WL, Scheer H, Yang X* & Zhao KH* The structure of allophycocyanin B from Synechocystis PCC 6803 reveals structural basis for extreme red-shift of terminal emitter in phycobilisomes Acta Cryst D 70:2558, doi:10.1107/S1399004714015776 (2014)
Munshi P, Snell EH, van der Woerd MJ, Judge RA, Myles DAA, Ren Z & Meilleur F* Neutron structure of the cyclic glucose-bound xylose isomerase E186Q mutant Acta Cryst D 70:414, doi:10.1107/S1399004713029684 (2014)
[2013]
Ren Z* Reverse engineering the cooperative machinery of human hemoglobin PLoS One 8:e77363, doi:10.1371/journal.pone.0077363 (2013)
Ren Z* Reaction trajectory revealed by a joint analysis of Protein Data Bank PLoS One 8:e77141, doi:10.1371/journal.pone.0077141 (2013)
Ren Z*, Chan P, Moffat K, Pai EF, Royer WE, Srajer V & Yang X Resolution of structural heterogeneity in dynamic crystallography Acta Cryst D 69:946, doi:10.1107/S0907444913003454 (2013)
[2010-2012]
Yang X* Structures of phytochromes GCK Roberts (ed) Encyclopedia of Biophysics Springer-Verlag Berlin Heilelberg (2012)
Ren Z*, Srajer V, Knapp JE & Royer WE Cooperative macromolecular device revealed by meta-analysis of static and time-resolved structures PNAS 109:107, doi:10.1073/pnas.1109213108 (2011)
Yang X*, Ren Z, Kuk J & Moffat K* Temperature-scan cryocrystallography reveals reaction intermediates in bacteriophytochrome Nature 479:428, doi:10.1038/nature10506 (2011)
Graber T*, Anderson S, Brewer H, Chen YS, Cho HS, Dashdorj N, Henning RW, Kosheleva I, Macha G, Meron M, Pahl R, Ren Z, Ruan S, Schotte F, Srajer V, Viccaro JP, Westferro F, Anfinrud P & Moffat K BioCARS: A synchrotron resource for time-resolved X-ray science J Synchrotron Rad 18:658, doi:10.1107/S0909049511009423 (2011)
Möglich A, Yang X, Ayers RA & Moffat K Structure and function of plant photoreceptors Annu Rev Plant Biol 61:21, doi:10.1146/annurev-arplant-042809-112259 (2010)
[2007-2009]
Yang X*, Kuk J & Moffat K* Conformational differences between the Pfr and Pr states in Pseudomonas aeruginosa bacteriophytochrome PNAS 106:15639, doi:10.1073/pnas.0902178106 (2009)
Yang X*, Kuk J & Moffat K* Crystal structure of Pseudomonas aeruginosa bacteriophytochrome: photoconversion and signal transduction PNAS 105:14715, doi:10.1073/pnas.0806718105 (2008)
Yang X, Stojkovic EA, Kuk J & Moffat K* Crystal structure of the chromophore binding domain of an unusual bacteriophytochrome, RpBphP3, reveals residues that modulate photoconversion PNAS 104:12571, doi:10.1073/pnas.0701737104 (2007)
Prior to 2007
Wu YL, Yang X, Ren Z, McDonnell DP, Norris JD, Willson TM & Greene GL* Structural basis for an unexpected mode of SERM-mediated ER antagonism Mol Cell 18:413, doi:10.1016/j.molcel.2005.04.014 (2005)
Schmidt M*, Pahl R, Srajer V, Anderson S, Ren Z, Ihee H, Rajagopal S & Moffat K Protein kinetics: Structures of intermediates and reaction mechanism from time-resolved X-ray data PNAS 101:4799, doi:10.1073/pnas.0305983101 (2004)
Krasilnikov A, Yang X, Pan T & Mondragon A* Crystal structure of the specificity domain of ribonuclease P Nature 421:760, doi:10.1038/nature01386 (2003)
Yang X, Gerczei T, Glover L & Correll C* Crystal structures of restrictocin-inhibitor complexes with implications for RNA recognition and base flipping Nature Struct Biol 8:968, doi:10.1038/nsb1101-968 (2001)
Correll CC, Munishkin A, Chan YL, Ren Z, Wool IG & Steitz TA* Crystal structure of the ribosomal RNA domain essential for binding elongation factors PNAS 95:13436, doi:10.1073/pnas.95.23.13436 (1998)
Perman B, Srajer V, Ren Z, Teng TY, Pradervand C, Ursby T, Bourgeois D, Schotte F, Wulff M, Kort R, Hellingwerf KJ & Moffat K* Energy transduction on the nanosecond time scale: early structural events in a xanthopsin photocycle Science 279:1946, doi:10.1126/science.279.5358.1946 (1998)
Yang X, Ren Z & Moffat K Structure refinement against synchrotron Laue data: strategies for data collection and reduction Acta Cryst D 54:367 doi:10.1107/S0907444997011517 (1998)
Genick UK, Borgstahl GEO, Ng K, Ren Z, Pradervand C, Burke PM, Srajer V, Teng TY, Schildkamp W, McRee DE, Moffat K & Getzoff ED Structure of a photocycle intermediate by millisecond time-resolved crystallography Science 275:1471, doi:10.1126/science.275.5305.1471 (1997)
Clifton IJ, Duke EMH, Wakatsuki S & Ren Z Evaluation of Laue diffraction patterns Meth Enzymol 277:448, doi:10.1016/S0076-6879(97)77025-3 (1997)
Srajer V, Teng TY, Ursby T, Pradervand C, Ren Z, Adachi SI, Schildkamp W, Bourgeois D, Wulff M & Moffat K Photolysis of the carbon monoxide complex of myoglobin: nanosecond time-resolved crystallography Science 274:1726, doi:10.1126/science.274.5293.1726 (1996)
Yang X* & Moffat K Insights into specificity of cleavage and mechanism of cell entry from the crystal structure of the highly specific Aspergillus ribotoxin, restrictocin Structure 4:837 doi:10.1016/S0969-2126(96)00090-1 (1996)
Ren Z*, Ng K, Borgstahl GEO, Getzoff ED & Moffat K Quantitative analysis of time-resolved Laue diffraction patterns J Appl Cryst 29:246 doi:10.1107/S0021889896000659 (1996)
Ren Z* & Moffat K Quantitative analysis of synchrotron Laue diffraction patterns in macromolecular crystallography J Appl Cryst 28:461 doi:10.1107/S0021889895003207 (1995)
Ren Z* & Moffat K Deconvolution of energy overlaps in Laue diffraction J Appl Cryst 28:482 doi:10.1107/S0021889895003219 (1995)
Ren Z, Meyer T & McRee DE* Atomic structure of a cytochrome c′ with an unusual ligand-controlled dimer dissociation at 1·8 Å resolution J Mol Biol 234:433 doi:10.1006/jmbi.1993.1597 (1993)